The ability to visualise and study objects at the nanoscale is important in many research fields especially in biology and medicine; this has been limited by the diffraction barrier of light in microscopy. For example, in a sample containing fluorophore molecules it can be difficult to distinguish between two closely spaced fluorophores due to the diffraction of light. The minimum distance that two fluorophores can have between them and still be resolved is determined by the Abbe limit.
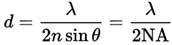
The Abbe limit depends on the wavelength of the light (λ) and the numerical aperture (NA) of the microscope. Typically, in fluorescence microscopy, point sources (or fluorophores) separated by less than 200 nm cannot be resolved. Therefore, two objects closer than 200 nm will look as if they are merged together.
Several microscopy techniques have been developed for single molecule detection, with a resolution beyond the diffraction limit. In 2014 Eric Betzig, Stefan Hell and William Moerner were awarded the Nobel Prize in Chemistry for their work in the development of super-resolved fluorescence microscopy, which enabled optical microscopy at the nanoscale.
Nanoscale studies of biological systems are essential to visualise enzyme systems, muscle proteins and cells functioning. Studying single molecule systems offer enormous potential for understanding the interactions, mechanisms and transport of individual biological macromolecules in the living cell.
The evolving post-genomic era of proteomics has raised the bar and requires the ability to track the behaviour, kinetics and mechanistic contribution of biochemically relevant single peptides, such as enzymes and carrier proteins. Moreover, special importance in biology is given to the direct, real-time visualisation of single biological macromolecules and their surroundings under in-vivo physiological conditions.
Over recent years, a family of super-resolution microscopy techniques have been developed for measurements below the diffraction limit. One of these techniques is super-resolution radial fluctuations (SRRF), developed by Ricardo Henriques and co-workers at University College London.
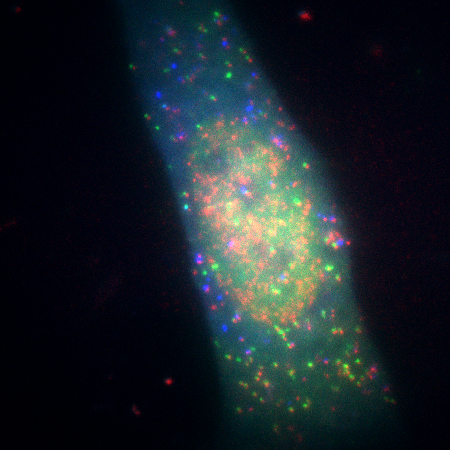
Single molecule imaging mRNA (red), during translation, and proteins, FLAGKDM5B (green) and HA-KDM5B (blue). Courtesy of Timothy J. Stasevich, IGAF, Colorado State University.
SRRF is a combination of temporal fluctuation analysis and localization microscopy. SRRF can be applied to data from imaging modes which include widefield, total internal reflectance microscopy (TIRF) and confocal, where short frame bursts (e.g. 50 frames) can be processed to deliver spatial resolution enhancements. Thus, SRRF can provide a route to super-resolution without the need for specialized optical hardware, exotic probes or very high-power densities.
SRRF is a post-processing super-resolution technique. Whereas, Andor’s implementation of SRRF as SRRF-Stream enables real time live cell super-resolution across a range of imaging modalities as opposed to post processing. SRRF-Stream is only avaliable on Andor's iXon Life and Ultra Series.
References
- Betzig, E., Patterson, G. H., Sougrat, R., Lindwasser, O. W., Olenych, S., Bonifacino, J. S., Davidson, M. W., Lippincott-Schwartz, J., Hess, H. F., Imaging Intracellular Fluorescent Proteins at Nanometer Resolution, Science, 2006, 313, 5793, 1642-1645, DOI: 10.1126/science.1127344
- Klar, T. A, Hell, S. W., Subdiffraction resolution in far-field fluorescence microscopy, 1999, Optics Letters. 24 (14): 954–956. DOI:10.1364/OL.24.000954.
- Gustafsson, N., Culley, S., Ashdown, G., Owen, D. M., Pereira, P. M., and Henriques, R. Fast live-cell conventional fluorophore nanoscopy with ImageJ through super-resolution radial fluctuations. Nature Communications 2016 7(12471):12471, DOI: 10.1038/ncomms12471


