The entire development of the nervous system can now be viewed with excellent temporal resolution using the exceptional capture rate of the Neo sCMOS combined with the Light Sheet Microscopy technique.
Philipp Keller and his colleagues at Janelia Farm Research Campus have developed one-photon and multi-photon SiMView technology and recorded cellular dynamics in entire Drosophila melanogaster embryos with 30-s temporal resolution throughout development. SiMView as its name implies is a light-sheet based one-photon and multi-photon simultaneous multiview imaging technique, which overcomes the limitations of sequential multiview strategies and enables quantitative systems-level imaging of fast dynamic events in large specimens. SiMView consists of a light sheet microscope with four synchronised optical arms, real-time electronics for long-term sCMOS-based image acquisition at 175 million voxels per second, and computational modules for high-throughput image registration, segmentation, tracking and real-time management of the terabytes of data recorded per specimen.
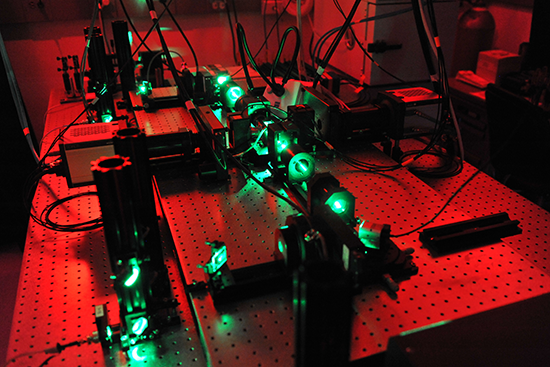

SiMView setup with 2 Neos
Keller’s main focus is to understand the developing nervous system. As a model system for the nervous system, Drosophila is an excellent candidate. For this study, transgenic embryos expressing elav(C155)-GAL4 and UAS-mCD8::GFP or Ftz-ng-GAL4,10XUAS-IVS-myr::GFP were used to visualize all postmitotic neurons and to perform time-lapse one-photon and multi-photon SiMView imaging of the entire embryonic nervous system to capture development of the central and peripheral nervous system and reveal fine details about the dynamics of axonal growth at excellent spatiotemporal resolution.
Keller’s SiMView imaging platform uses an orthogonal arrangement of four independently operated optical arms. One pair of the optical arms is used for bidirectional light-sheet illumination with two long-working distance objectives. The other pair, arranged at a right angle to the first, is used for bidirectional fluorescence detection with high-numerical-aperture objectives. In this set-up, the sample is located in a 25-ml chamber with full optical access from all sides to enable simultaneous light-sheet illumination and fluorescence detection with four objectives. By splitting the microscope into two horizontal layers, full optical access and maximum specimen positioning control are enabled. All optical systems are located in the upper layer, whereas the specimen positioning system is upright, in contrast to previous light-sheet microscopes. The benefit of this is that the sample can be embedded in exceptionally soft (0.4 %) agarose gel, which allows long-term imaging of entire developing Drosophila and zebrafish embryos under physiological conditions.
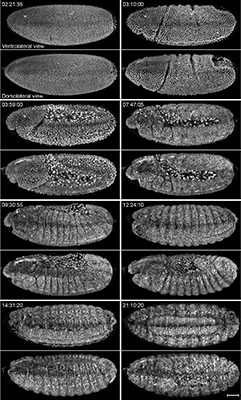
Different views of the embryo during the developmental stages
In order to acquire the millions of high-speed and high resolution images with two Neo sCMOS cameras simultaneously over the entire development of the Drosophila embryo, Keller and his team control the entire SiMView system on a high-performance real-time controller. The controller communicates with a host computer workstation to coordinate the simultaneous acquisition at a rate of 350 MB s-1. This control framework performs the alignment of the microscope’s four optical arms with submicron precision directly on each specimen, using automated software modules for auto-focussing and light-sheet positioning.
Due to the amount of data produced when performing light sheet microscopy on entire embryos, Keller designed the SiMView computational backbone for high-speed imaging experiments with up to several days of continuous image acquisition. This system is capable of recording more than 1 million high-resolution images in uninterrupted high-speed imaging sessions with a total data set size of 10 TB per specimen. In addition to this, Keller’s group have a secondary 10-gigabit s-1 glass fibre network-based acquisition pipeline which allows recording up to 10 million high-resolution images or 100 TB per specimen for long-term imaging sessions. The SiMView real-time data compaction module for lossless 3-D wavelet compression allows for the maximum recording capacity of 1 petabyte.
From comparative analysis of the images acquired with SiMView, single view and sequential multiview, Keller concluded that SiMView imaging improves coverage over single view imaging by almost fourfold and reveals cellular dynamics in the entire early embryo. Furthermore, SiMView imaging shows several advantages over sequential multiview imaging: it eliminates temporal and spatial fusion artifacts during fast cellular movements, improves temporal resolution and provides quantitative data for subsequent computational image analyses such as automated cell tracking. Importantly, SiMView reduces the photobleaching and phototoxic effects on the specimen itself by avoiding the continual acquisition of overlapping regions required for sequential multiview image registration. In addition, SiMView reveals detailed information at the sub-cellular level which in sequential multiview imaging is misrepresented. Moreover, SiMView accurately records fast local nuclear movements, whereas sequential imaging often leads to duplicate structures and distorted nuclear shapes.
One-photon SiMView provided superior signal-to-noise ratio for small penetration depths but failed to capture structures deep inside the embryo. In contrast, two-photon SiMView reduced autofluorescence compared and provided near complete physical coverage of the embryo even for late development stages.
According to Keller the SiMView imaging technique opens doors to high-throughput high-content screening, fast functional imaging and comprehensive quantitative analyses of cellular dynamics in entire developing organisms.
Andor Solutions for light sheet microscopy
Andor' sCMOS cameras are ideal detectors for high-speed light sheet microscopy. Neo 5.5, Zyla 5.5 and Zyla 4.2 all offer a large field of view and high resolution without compromising read noise or frame rate, and the 6.5 μm pixel size is ideally suited to light sheet microscopy. The small pixel size of 6.5 micrometers ensures sufficient oversampling of the pointspread-function, even for objectives with low magnification (e.g. 10x NA 0.3; 20x NA 0.5). The read noise is very low, even when compared with the highest-performance slow readout CCDs. Zyla 5.5 can achieve a read noise down to 1.45 e– rms, while Zyla 4.2 has a read noise of 1.1 e- rms, both without signal amplification, while reading out 5.5 and 4.2 megapixels at 100 frames per second respectively.



